Estimates a multidimensional regression function (and its first derivatives) using local polynomial kernel smoothing (and linear binning).
locpol(x, ...)
# Default S3 method
locpol(
x,
y,
h = NULL,
nbin = NULL,
type = c("linear", "simple"),
degree = 1 + as.numeric(drv),
drv = FALSE,
hat.bin = FALSE,
ncv = 0,
set.NA = FALSE,
...
)
# S3 method for class 'bin.data'
locpol(
x,
h = NULL,
degree = 1 + as.numeric(drv),
drv = FALSE,
hat.bin = FALSE,
ncv = 0,
...
)
# S3 method for class 'svar.bin'
locpol(x, h = NULL, degree = 1, drv = FALSE, hat.bin = TRUE, ncv = 0, ...)
# S3 method for class 'bin.den'
locpol(x, h = NULL, degree = 1 + as.numeric(drv), drv = FALSE, ncv = 0, ...)
locpolhcv(
x,
y,
nbin = NULL,
objective = c("CV", "GCV", "MASE"),
degree = 1 + as.numeric(drv),
drv = FALSE,
hat.bin = FALSE,
set.NA = FALSE,
ncv = ifelse(objective == "CV", 2, 0),
cov.dat = NULL,
...
)Arguments
- x
a (data) object used to select a method.
- ...
further arguments passed to or from other methods (e.g. to
hcv.data).- y
vector of data (response variable).
- h
(full) bandwidth matrix (controls the degree of smoothing; only the upper triangular part of h is used).
- nbin
vector with the number of bins on each dimension.
- type
character, binning method:
"linear"(default) or"simple".- degree
degree of the local polynomial used. Defaults to 1 (local linear estimation).
- drv
logical; if
TRUE, the matrix of estimated first derivatives is returned.- hat.bin
logical; if
TRUE, the hat matrix of the binned data is returned.- ncv
integer; determines the number of cells leaved out in each dimension. Defaults to 0 (the full data is used) and it is not normally changed by the user in this setting. See "Details" below.
- set.NA
logical. If
TRUE, sets the bin averages corresponding to cells without data toNA.- objective
character; optimal criterion to be used ("CV", "GCV" or "MASE").
- cov.dat
covariance matrix of the data or semivariogram model (of class extending
svarmod). Defaults to the identity matrix (uncorrelated data).
Value
Returns an S3 object of class locpol.bin (locpol + bin data + grid par.).
A bin.data object with the additional (some optional) 3 components:
- est
vector or array (dimension
nbin) with the local polynomial estimates.- locpol
a list with 7 components:
degreedegree of the polinomial.hbandwidth matrix.rmresidual mean.rsssum of squared residuals.ncvnumber of cells ignored in each direction.hat(if requested) hat matrix of the binned data.nrl0(if appropriate) number of cells with data (binw > 0) and missing estimate (est == NA).
- deriv
(if requested) matrix of first derivatives.
locpol.svar.bin returns an S3 object of class np.svar
(locpol semivar + bin semivar + grid par.).
locpol.bin.den returns an S3 object of class np.den
(locpol den + bin den + grid par.).
Details
Standard generic function with a default method (interface to the
fortran routine lp_raw), in which argument x
is a vector or matrix of covariates (e.g. spatial coordinates).
If parameter nbin is not specified is set to pmax(25, rule.binning(x)).
A multiplicative triweight kernel is used to compute the weights.
If ncv > 0, estimates are computed by leaving out cells with indexes within
the intervals \([x_i - ncv + 1, x_i + ncv - 1]\), at each dimension i, where \(x\)
denotes the index of the estimation position. \(ncv = 1\) corresponds with
traditional cross-validation and \(ncv > 1\) with modified CV
(see e.g. Chu and Marron, 1991, for the one dimensional case).
Setting set.NA = TRUE (equivalent to biny[binw == 0] <- NA)
may be useful for plotting the binned averages $biny
(the hat matrix should be handled with care).
locpolhcv calls hcv.data to obtain an "optimal"
bandwith (additional arguments ... are passed to this function).
Argument ncv is only used here at the bandwith
selection stage (estimation is done with all the data).
References
Chu, C.K. and Marron, J.S. (1991) Comparison of Two Bandwidth Selectors with Dependent Errors. The Annals of Statistics, 19, 1906-1918.
Rupert D. and Wand M.P. (1994) Multivariate locally weighted least squares regression. The Annals of Statistics, 22, 1346-1370.
See also
Examples
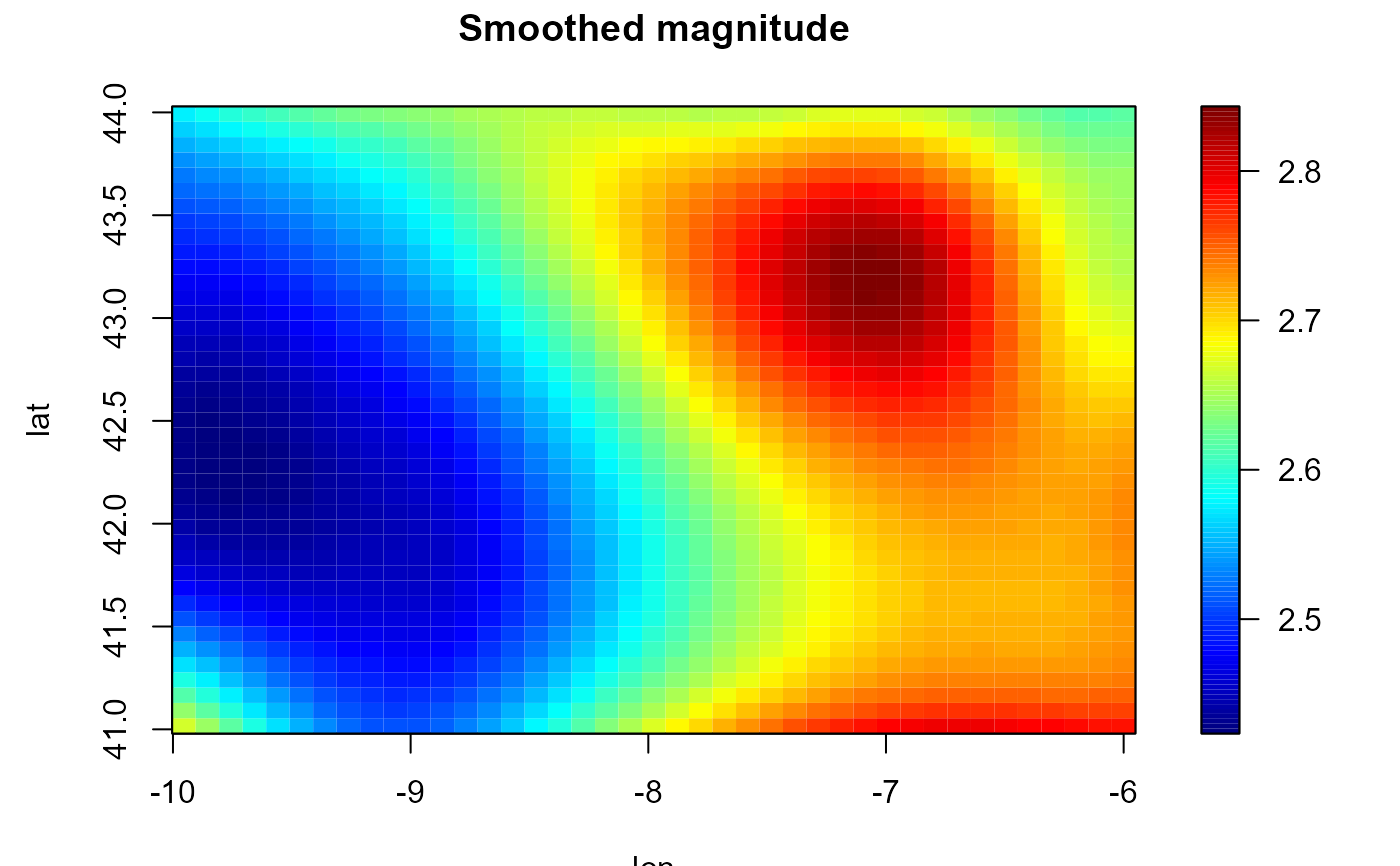
lp <- locpol(earthquakes[, c("lon", "lat")], earthquakes$mag, h = diag(2, 2), nbin = c(41,41))
simage(lp, main = "Smoothed magnitude", reset = FALSE)
contour(lp, add = TRUE)
 bin <- binning(earthquakes[, c("lon", "lat")], earthquakes$mag, nbin = c(41,41))
lp2 <- locpol(bin, h = diag(2, 2))
all.equal(lp, lp2)
#> [1] TRUE
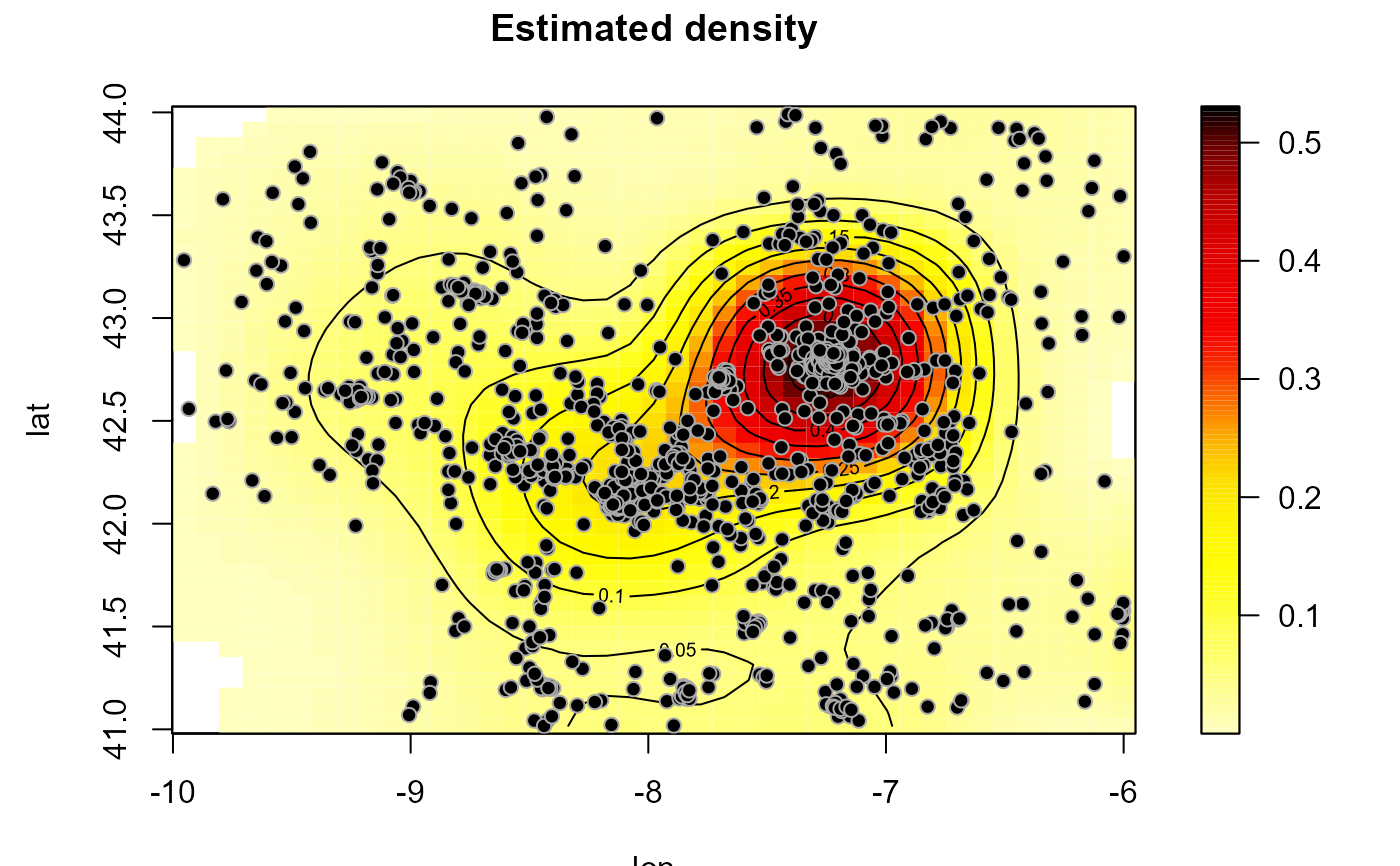
den <- locpol(as.bin.den(bin), h = diag(1, 2))
plot(den, log = FALSE, main = 'Estimated density')
bin <- binning(earthquakes[, c("lon", "lat")], earthquakes$mag, nbin = c(41,41))
lp2 <- locpol(bin, h = diag(2, 2))
all.equal(lp, lp2)
#> [1] TRUE
den <- locpol(as.bin.den(bin), h = diag(1, 2))
plot(den, log = FALSE, main = 'Estimated density')